|
The battle of our immune system with that of pathogens has been going on for millennia (longer than battle of Avengers and all their their nemesis combined!). The bugle of battle was blown with the occurrence of first multicellular organism 3.5 million years ago and with the rise of first parasite, who knows that it might be our own mitochondria? Its a speculation though, but lack of explanation for its origin and its astounding similarity with prokaryotes, makes it open to any body's guess. However, in these millions of years both our immune system and pathogens evolved playing the game of hide and seek and developed several weapons in their armory to outsmart each other. Genomic analysis of plants and animals provides evidence that a sophisticated mechanism of host defense was in existence by the time the ancestors of plants and animals diverged. This system, is being shared by plants and animals, and the Toll pathway of NFκB activation is an example, demonstrated conclusively in fruit flies such as Drosophila and in vertebrates such as mice and humans and also believed to occur in plants in the form of Leucin Rich Repeats (LRRs). Now let us introduce ourselves with the three super hero of our immune system. 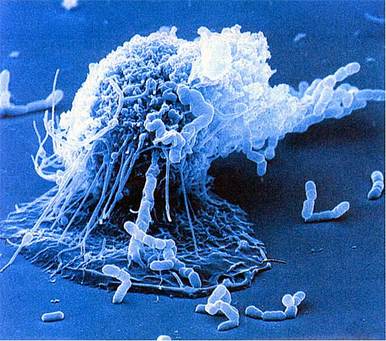
Macrophage
Granulocyte-Monocyte progenitor cells in the bone marrow differentiate into pro-monocytes, which upon entering blood, further differentiates into mature monocytes. Monocytes circulate in the bloodstream for about 8 h, during which they migrate into the tissues and differentiate into specific tissue macrophages. Enlarge five- to ten fold; its intracellular organelles increase in both number and complexity; and it acquires increased phagocytic ability, produces higher levels of hydrolytic enzymes, and begins to secrete a variety of soluble factors.(Remember Hulk!). Whenever I think of macrophage it sounds to me like Hulk, and I have reasons to backup the claim, first it is one of the biggest cell observable under microscope with size approx 21 μm (micrometres). Hulk likes to smash, and our macrophage likes to phagocytose its oponents. Macrophages are capable of ingesting and digesting exogenous antigens, such as whole microorganisms and insoluble particles, and endogenous matter, such as injured or dead host cells, cellular debris, and activated clotting factors. Moreover years of selection pressure has made macrophage a more leathal enemy as it equipped itself with many more weapons, such as Opsonization, production of reactive oxygen intermediates (ROIs) and reactive nitrogen intermediates that have potent antimicrobial properties (consider WMDs), along with a group of antimicrobial and cytotoxic peptides, commonly known as defensins. Defensin peptides have been shown to form ion-permeable channels (pores!) in bacterial cell membranes, and can kill a variety of bacteria, including Staphylococcus aureus, Streptococcus pneumoniae, Escherichia coli,Pseudomonas aeruginosa, and Haemophilus influenzae. Consider Hulk with a gun huh! 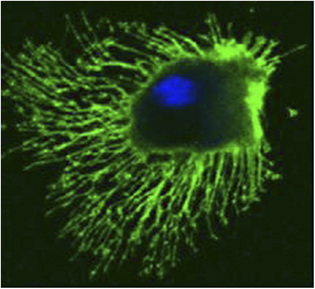
Dendritic Cells (DCs)
Dendritic cells are derived from hematopoietic bone marrow progenitor cells, these progenitor cells initially transform into immature dendritic cells. Dcs acquired its name because it is covered with long membrane extensions that resemble the dendrites of nerve cells. DCs constitutively express high levels of both class II MHC molecules and members of the co-stimulatory B7 family. For this reason, they are more potent antigen-presenting cells than macrophages and B cells, both of which need to be activated before they can function as antigen-presenting cells (APCs). The dendritic cells are constantly in communication with other cells in the body. This communication can take the form of direct cell–cell contact based on the interaction of cell-surface proteins. An example of this includes the interaction of the membrane proteins of the B7 family of the dendritic cell with CD28 present on the lymphocyte. However, the cell–cell interaction can also take place at a distance via cytokines. Following microbial invasion or during inflammation, mature and immature forms of Langerhans cells and interstitial dendritic cells migrate into draining lymph nodes, where they make the critical presentation of antigen to TH cells that is required for the initiation of responses by those key cells. Looks like Captain America isn't it? Flexible, resourceful, communicating crucial intel to raise deffence, planning, integrating and keeps the team going. 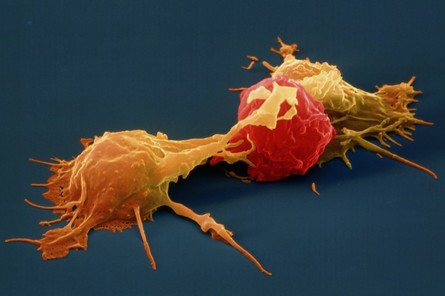
Natural Killer Cell (NK cell)
Natural Killer Cell (NK cell)consists of a small population of large, granular lymphocytes that display cytotoxic activity and are analogous to that of cytotoxic T cells. Cytotoxic activity is displayed against a wide range of cells, both viral infected and transformed. If nature could issue a license to kill NK cells would be the best candidate for it because unlike cytotoxic T cells, NK cell can directly induce the death of tumor cells and virus-infected cells in the absence of specific immunization. Armed with an array of receptors that can either stimulate NK cell reactivity (activating receptors eg. NKG2) or dampen NK cell reactivity (inhibitory receptors e.g. KIRs), NK cells are smart like Tony Stark and lethal like Iron Man! However do not think that all this fire power is uncontrolled, it has a very smart control system like JARVIS, which avoids auto-reactivity, by initiating an education system where NK cells acquire self-tolerance. Unlike T-cells however the potentially autoreactive NK cells are not generally clonally deleted but instead a maintained in a state of hyporesponsiveness or anergy. Several findings suggest that the responsiveness of mature NK cells is not fixed but may adapt to a changing environment in vivo. It is observed that persistent stimulation without inhibition results in NK cell hyporesponsiveness, whereas persistent stimulation coupled with commensurate inhibition results in NK cell responsiveness. These results suggest that NK cell tuning might occur throughout the lifetime of the NK cell under steady-state conditions. In infected animals, however, hyporesponsive NK cells are converted to a higher state of responsiveness. Reference:
0 Comments
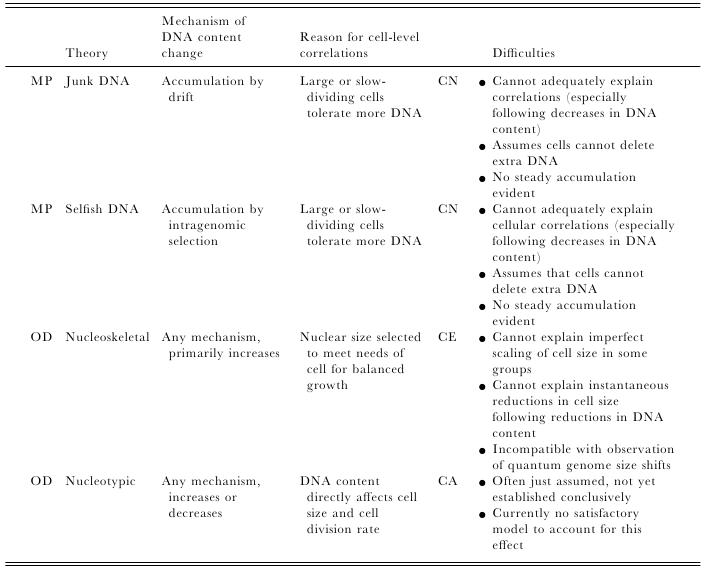
Most of us grow up listening, reading and learning the fact that we "Human" beings can boast of being the most evolved and a higher organism. Earlier than 1960, the image below would have been correct and sensible. However our idea of supremacy in terms of genome size and number of genes takes a flak when we first started looking at the complexity of genome size, it was soon realized that the large genomes were often composed of huge chunks of repetitive DNA, while only a few percent of the genome in these organisms were unique. Now lets have a look at the past and try to figure out the origin of this concept. 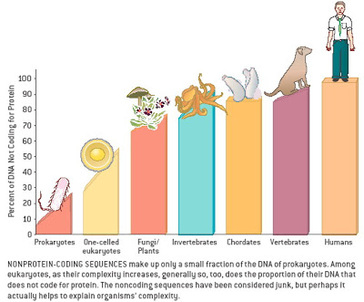 Classically biologists recognize that the living world comprises two types of organisms. Prokaryotes and Eukaryotes. Assuming that you already know what prokaryotes and eukaryotes are, I am not going to dive into the difference between the two. So what is C-value? 'C-value ', of an organism is defined as the total amount of DNA contained within its haploid chromosome set. Prokaryotic cells typically have genomes smaller than 10 megabases (Mb), while the genome of single cell eukaryote is typically less than 50Mb. Therefore for simplicity's sake here we are not comparing the genomes from two classes of organisms together. However eukaryotes alone show immense diversity among their genome sizes, from the smallest eukaryote being less than 10 Mb in length, and the largest over 100 000 Mb, and all these observation seems coinciding to a certain extent with the complexity of the organism, the simplest eukaryotes such as fungi having the smallest genomes, and higher eukaryotes such as vertebrates and flowering plants having the largest ones. So it seems fair to think that, complexity of an organism is related to the number of genes in its genome - higher eukaryotes need larger genomes to accommodate the extra genes. However, in fact this correlation is far from being precise: if it were, then the nuclear genome of the yeast S. cerevisiae, which at 12 Mb is 0.004 times the size of the human nuclear genome, would be expected to contain 0.004 × 35 000 genes, which is just 140. In fact the S. cerevisiae genome contains about 5800 genes! Therefore for many years this lack of precise correlation between the complexity of an organism and the size of its genome was looked on as a bit of a puzzle, and called as C-value paradox/C-value enigma. Questions raised by C-value paradox The C-value paradox not only represents one question, but it rather raises three of them, as suggested by T. R. Gregory(2007), 1) the generation of large-scale variation in genome size, which may occur by continuous or quantum processes, (2) the non-random distributions of genome size variation, whereby some groups vary greatly and others appear constrained, and (3) the strong positive relationship between C-value and nuclear and cell sizes and the negative correlation with cell division rates. Therefore any proposed solution must try to solve these three problems as well. Now we biologists are pretty good at dividing ourselves among different school of thoughts (remember the RNA and DNA world!) and the C-value paradox wasn't an exception either. Nonetheless two school of thoughts emerged here too, one proposing Mutation pressure theories and the other proposing Optimal DNA theories. The table shown below summarizes the theories proposed along with their proposed mechanism. Each theory can be classified according to its explanation for the accumulation or maintenance of DNA (MP,mutation pressure theory ; OD, optimal DNA theory) and according to its explanation for the observed cellular correlations (CN, coincidental, CE, coevolutionary, CA, causative). Note that these theories are not necessarily mutually exclusive in all respects, since the optimal DNA theories do not specify the mechanism(s) of DNA content change and can include those presented by both mutation pressure theories. (ref: GREGORY, T. R. (2001),page # 69) So what is the most plausible explanation of C-value paradox? In 1980s, two landmark papers, by Orgel and Crick and by Doolittle and Sapienza, established a strong case against 'selfish DNA elements' which we better know as transposons. They proposed that ‘selfish DNA’ elements, such as transposons, essentially act as molecular parasites, replicating and increasing their numbers at the (usually slight) expense of a host genome, i.e these elements functions for themselves while providing little or no selective advantage to the host. Computational genomic studies have shown that transposable elements invade in waves over evolutionary time, sweeping into a genome in large numbers, then dying and decaying away leaving the 'Junk DNA' in its trail. 45% of the human genome is detectably derived from these transposable elements. Therefore we can say that C-value paradox is mostly (though not entirely) explained by different loads of leftovers from transposable elements and larger the genomes longer is the trail leftover by transposons. So, if the C-value paradox is explained and rested for good, why dig it up again? Recent publications discussing the outcome of ENCODE (Encyclopedia Of DNA Elements) project suggest that 80% of the human genome is reproducibly transcribed, bound to proteins, or has its chromatin specifically modified. Moreover the previously considered junk DNA is found to be biochemically active disapproving the 'Junk DNA' theory. Now in the light of ENCODE data it is pertinent that scientists need to come up with a alternative hypothesis capable of explaining C-value paradox, for mutational load, and for how a large fraction of eukaryotic genomes is composed of neutrally drifting transposon-derived sequences. Reference:
The Toll family of receptors
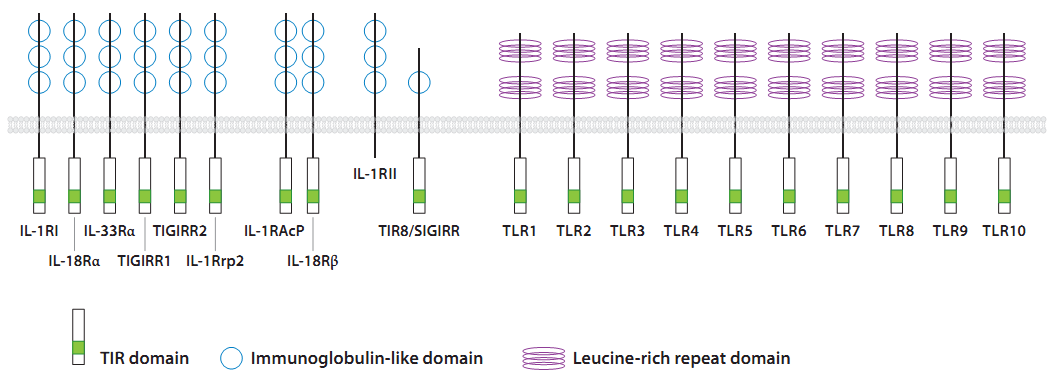
Toll Like Receptors or TLRs are type I transmembrane proteins of the Interleukin-1 receptor (IL-1R) family that possess an N-terminal leucine-rich repeat (LRR) domain for ligand binding, a single transmembrane domain, and a C-terminal intracellular signaling domain. The TLR C terminus is homologous to the intracellular domain of the IL-1R and is thus referred to as the Toll/IL-1 receptor (TIR) domain. TLRs are expressed at the cell membrane and in sub cellular compartments such as the endosomes and are widely expressed in many cell types, including non hematopoietic epithelial, endothelial cells. Although most cell types express only a select subset of these receptors. However hematopoietically derived sentinel cells, such as macrophages, neutrophils, and dendritic cells (DCs), express most of the TLRs, with some variation in different subsets, e.g., between conventional DCs and plasmacytoid DCs. Thus far, 13 mammalian TLRs, 10 in humans and 13 in mice, have been identified (Beutler 2004). TLRs 1– 9 are conserved among humans and mice, yet TLR10 is present only in humans and TLR11 is functional only in mice. Although much is known about the ligands and signaling pathways of TLRs 1–9 and 11, the biological roles of TLRs 10, 12, and 13 remain unclear, as their expression patterns, ligands, and modes of signaling have yet to be defined.
TLRs mediate initial responses in innate immunity and are required for the development of the adaptive immune response. Toll-like receptors (TLRs) enable innate immune recognition of endogenous and exogenous prototypic ligands. They also orchestrate innate and adaptive immune response to infection, inflammation, and tissue injury. Given their significance in the immune response, it is not surprising that genetic variations of TLRs can affect their function and by extension affect the response of the organism to environmental stimuli. The genetics of TLRs provides important insights in gene-environment interactions in health and disease, and it may enable scientists to assess patients’ susceptibility to diseases or predict their response to treatments. Evolutionary genetics of TLRs In the domain of the evolutionary genetics of infectious diseases, the aim is to identify the evolutionary footprints of natural selection exerted by past infections in the genome of present day healthy human populations. Given the tremendous selective pressure that pathogens have exerted in the past, and continue to exert, it is hardly surprising that some of the strongest evidence for selection, of various types and intensities, in the human genome has actually been obtained for genes involved in immunity or host defense as immunity related functions seem to be a privileged target of natural selection in the human species as a whole (with respect to other primates) and in different human populations from diverse geographic regions. Phylogenetic studies have indicated an ancient origin for TLR genes, some 700 million years ago, suggesting that TLR-mediated immune responses originated in the common ancestor of bilaterian animals. However, several recent, independent lines of evidence genomic, phylogenetic, and functional data have suggested that the similarities and differences between TLR-mediated innate immunity functions in insects and vertebrates may instead have resulted from convergent evolution. Convergent evolution refers the phenomenon where organisms that are not closely related independently evolve similar traits as a result of having to adapt to similar environments or ecological niches. Another study showed that vertebrate TLRs can be divided into six major families, with all the TLRs within a given family recognizing the same general or specific class of microbial compound. The patterns of interspecies divergence and levels of polymorphism in various primates, including humans, have recently been investigated. Signatures of accelerated evolution (species-wide positive selection) was found across primate species for most TLRs, with the strongest evidence of this obtained for TLR1 and TLR4, which have been independently targeted by positive selection. However, within each primate species, the patterns of nucleotide variation were generally constrained.
TLRs and Diseases TLRs In Pulmonary Diseases Current data suggest that TLR signaling can modify both allergic asthma and chronic obstructive pulmonary disease (COPD). Activation of TLRs can be either beneficial or detrimental depending on many host factors, as well as dose, duration, and intensity of expo-sure to TLR ligands. Multiple epidemiologic studies have associated childhood exposure to TLR ligands with protection against develop-ing allergic asthma later in life (the “hygiene hypothesis”); for example, individuals living on farms have a reduced risk of developing hay fever or asthma. The most extensively studied TLR is TLR4. A study of asthma specifically associated with LPS in house dust showed that people with the TLR4 polymorphism Asp299Gly had a de-creased risk of bronchoreactivity. These observations are consistent with the hypothesis that LPS can exacerbate existing airway inflammation and that individuals with the Asp299Gly polymorphism have diminished pulmonary responses to LPS. TLRs In CardioVascular Disease Atherosclerosis is an inflammatory process, and innate immunity has been shown to par-ticipate in the development and rupture of atherosclerotic plaques. TLR4 polymorphisms that render the receptor less responsive to its ligands would therefore be expected to hinder the development and progression of atherosclerosis. Indeed, the Asp299Gly poly-morphism has been associated with decreased atherosclerosis, decreased risk for acute coronary events , and an improved re-sponse to statin treatment. The exact mechanism of this beneficial effect is un-known; however, TLRs are expressed on several cells that participate in the atherosclerotic plaque, such as macrophages, dendritic cells, endothelia, smooth muscle cells, and lymphocytes. As the involvement of innate immunity in atherosclerosis is better understood, more genetic factors are likely to be discovered to influence both the suscep-tibility to cardiovascular disease and the response to treatment. TLRs In Cancer In Cancer inflammation acts as a double-edged sword. On one hand, chronic inflammation is associated with carcinogenesis, and cancer is a complication of chronic inflammatory conditions such as Crohn’s disease, chronic cystitis, and hepatitis. TLR activation leads to production of NF-κ B, which is associated with carcinogenesis and chemoresistance On the other hand, the immune system is necessary for the elimination of malignant cells, and immunosuppressed patients are a risk for the development of cancer. Immunotherapy (i.e., use of the patient’s own immune system to combat cancer, with the aid of vaccination or adjuvants) has been gaining attention as a potential treatment option in cancer. Indeed, we are only now beginning to understand how cancer cells evade elimination by modifying the innate and adaptive immune responses of the host. Activation of the immune system may therefore be a viable approach to cancer therapy. Reference: 1 . Casanova, Jean-Laurent, Laurent Abel, and Lluis Quintana-Murci. "Human TLRs and IL-1Rs in host defense: natural insights from evolutionary, epidemiological, and clinical genetics." Annual review of immunology 29 (2011): 447-491. 2. Pancer, Zeev, and Max D. Cooper. "The evolution of adaptive immunity." Annu. Rev. Immunol. 24 (2006): 497-518. 3. Garantziotis, Stavros, et al. "The Effect of Toll-Like Receptors and Toll-Like Receptor 4. Roach, Jared C., et al. "The evolution of vertebrate Toll-like receptors." Proceedings of the National Academy of Sciences of the United States of America 102.27 (2005): 9577-9582. 5. CE Sawian, SD Lourembam, A Banerjee, S Baruah “ Polymorphisms and expression of TLR4 and 9 in malaria in two ethnic groups of Assam, northeast India” Innate immunity 19 (2), 174-183 |
AuthorHello! My name is Arunabha Banerjee, and I am the mind behind Biologiks. Leaning new things and teaching biology are my hobbies and passion, it is a continuous journey, and I welcome you all to join with me Archives
June 2024
Categories
All
|

 RSS Feed
RSS Feed



